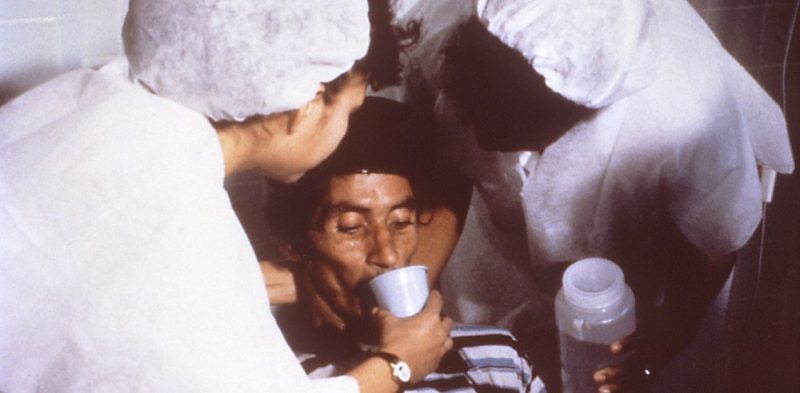
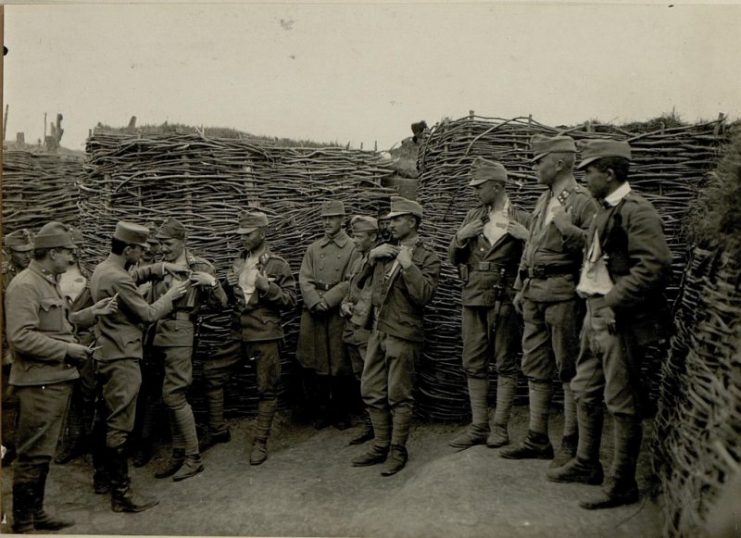
During the trench fighting in the First World War, the world experienced a global cholera pandemic, now known as the sixth pandemic.
Professor Nick Thomson, from the Wellcome Sanger Institute and his team of researchers, have mapped the genetic code of the oldest publicly available cholera-causing strain of bacteria. The strain, Vibrio cholerae, has been stored for over 100 years since it was isolated from a British soldier who had cholera-like symptoms during the First World War.
Cholera can be a deadly disease. The disease causes severe diarrhea and is caused by a bacterium, toxigenic V. cholerae, that is found in contaminated food and water. Cholera is prevalent after natural disasters and spreads like wildfire. It has been the cause of many pandemics, and thousands of deaths around the world.
Professor Thompson’s report reveals that the strain investigated is unique and a distant and non-toxigenic strain of V. cholerae that causes so many illnesses.
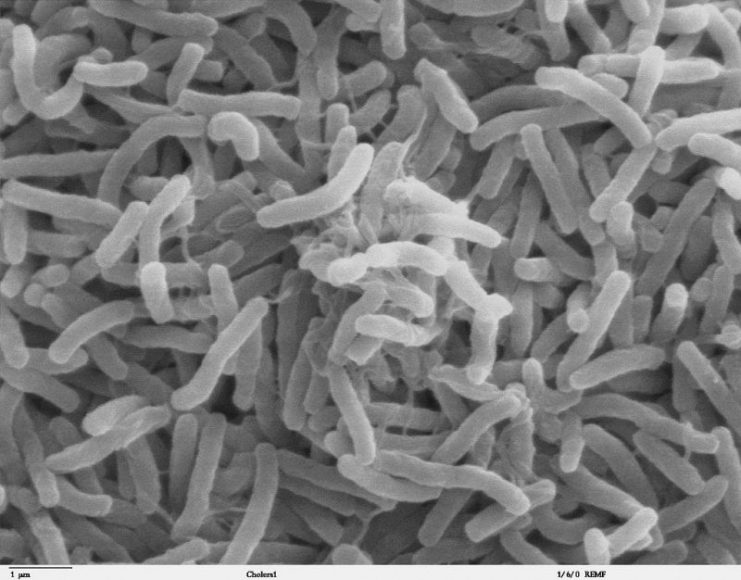
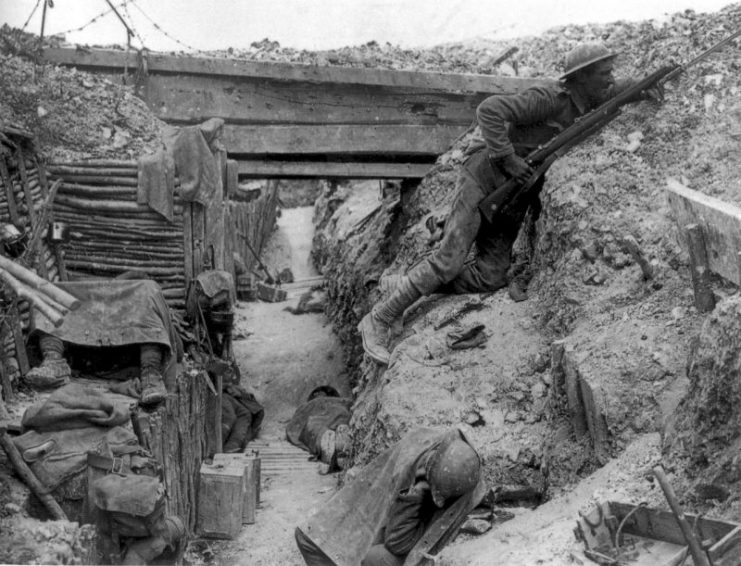
During the trench fighting in the First World War, the world experienced a global cholera pandemic, now known as the sixth pandemic. However, hardly any of the soldiers serving with the British Expeditionary Forces fell ill with cholera. The threat of cholera was taken very seriously during the war, but fortunately those fears did not come to pass.
The particular sample used by the research team was taken from a stool sample given by a soldier who was recovering from what was termed ‘choleraic diarrhea’ in Egypt in 1916. Since 1920, the extracted bacterium has been stored in the National Collection of Type Cultures (NCTC).

The research team resurrected the isolated bacterium and sequenced its genome. The sequencing results showed that the bacterium was a strain of V. cholerae, but it was incapable of causing the type of cholera responsible for pandemics. Further, it was unrelated to the strain of cholera that caused the sixth pandemic that was so prevalent during the war.
Professor Thomson spoke about the work done to decode the genome of what is thought to be the oldest stored live sample of V. cholerae. The importance of this work cannot be underestimated, since studying bacteria strains from different time periods provides insight into the evolution of this species of bacteria and helps in understanding the links between disease outbreaks and human history.
The fact that this specific bacterium did not cause an outbreak of cholera does not mean studying it is not essential. This slice of history is significant as it is part of the history of cholera and adds to our understanding of the disease.
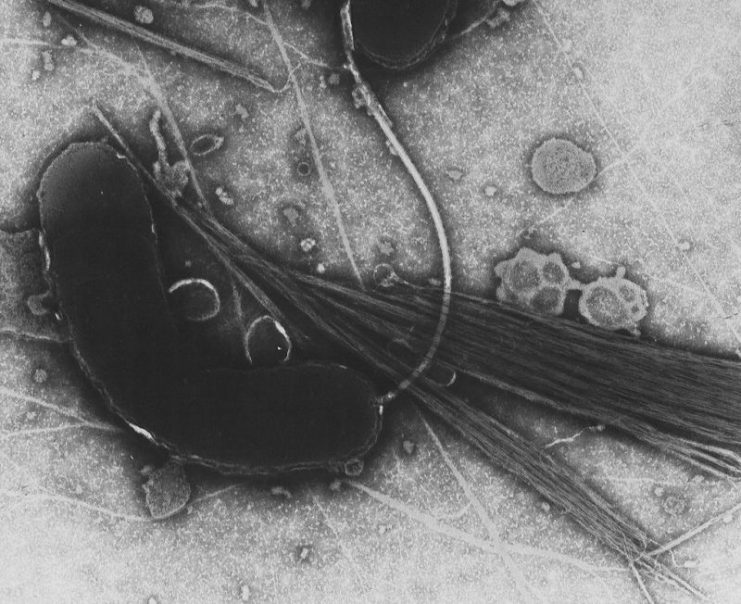
Another member of the research team, Matthew Dorman said that literature research showed that there was something unusual about this particular bacterium. Looking at the bacterium under a microscope, they saw that it lacked a flagellum or tail and was therefore unable to swim. They then found that that there was a mutation in a gene that is critical for the development of a flagella, which may explain why the bacterium had no tail.
https://youtu.be/tx41dFGvurQ
They were delighted to see that data from their genome project concurred with the historical literature. Reports from the time the sample was taken indicate that the soldier suffered from diarrhea, similar to what is seen in cholera cases. However, the recent study showed that he was not suffering from cholera since he was infected with a non-toxigenic bacterium.

Another unexpected finding was that the bacterium had an ampicillin resistant gene. This is another piece of evidence suggesting antibiotic resistance was in existence before the development of antibiotics. The researchers believe that this gene developed in response to naturally occurring antibiotics.
Read another story from us: WWI And The Paranormal: Spooky Tales From The Trenches
The head of the NCTC said that they had collected, maintained, and grown a collection of over 5,000 strains of bacteria, some of which date back over a hundred years. Studying these bacteria provides valuable data on the bacterial evolution and their influence on human history.